Analyzing an Off-flavor Component of Adhesive using Group Analysis Function of msFineAnalysis AI
MSTips No. 419
Introduction
Electron ionization (EI) is one of the most popular ionization methods used in gas chromatography-mass spectrometry (GC-MS). Consequently, compounds are typically identified by a mass spectral database search using EI mass spectra. Because molecular ions are often weak or absent in 70 eV EI mass spectra, identification of unknowns can be difficult by EI alone. In these cases, soft ionization (SI) can be very helpful for producing and identifying molecular ions. Recently, JEOL began developing an integrated qualitative analysis workflow that automatically combines and interprets the information from EI and SI data1). And then in 2018, we introduced our integrated qualitative analysis software "msFineAnalysis" which uses both EI and SI data to improve compound identification for GC-MS applications. Despite the fact that msFineAnalysis was automatically able to determine the molecular formula and partial structure information from EI fragment ion formulas, the actual structural formulas still required manual analysis using chemical compositions. To address this, we then developed an automated structure analysis software package entitled "msFineAnalysis AI" which uses artificial intelligence (AI) to predict EI mass spectra from chemical structures2).
The msFineAnalysis AI software has a Group analysis function that enables easy extraction of specific compounds. In MSTips No. 417, we introduced an overview of group analysis function and example of its application for vinyl acetate resin samples. The group analysis function includes a list of characteristic fragment ions and neutral losses as described in MSTips No. 417, as well as an "additive list" containing major additives for polymeric materials and an "off-flavor list" containing off-flavor components. In this MSTips, we will introduce off-flavor compounds analysis results using this group analysis function with "off-flavor list".
Experimental

msFineAnalysis AI
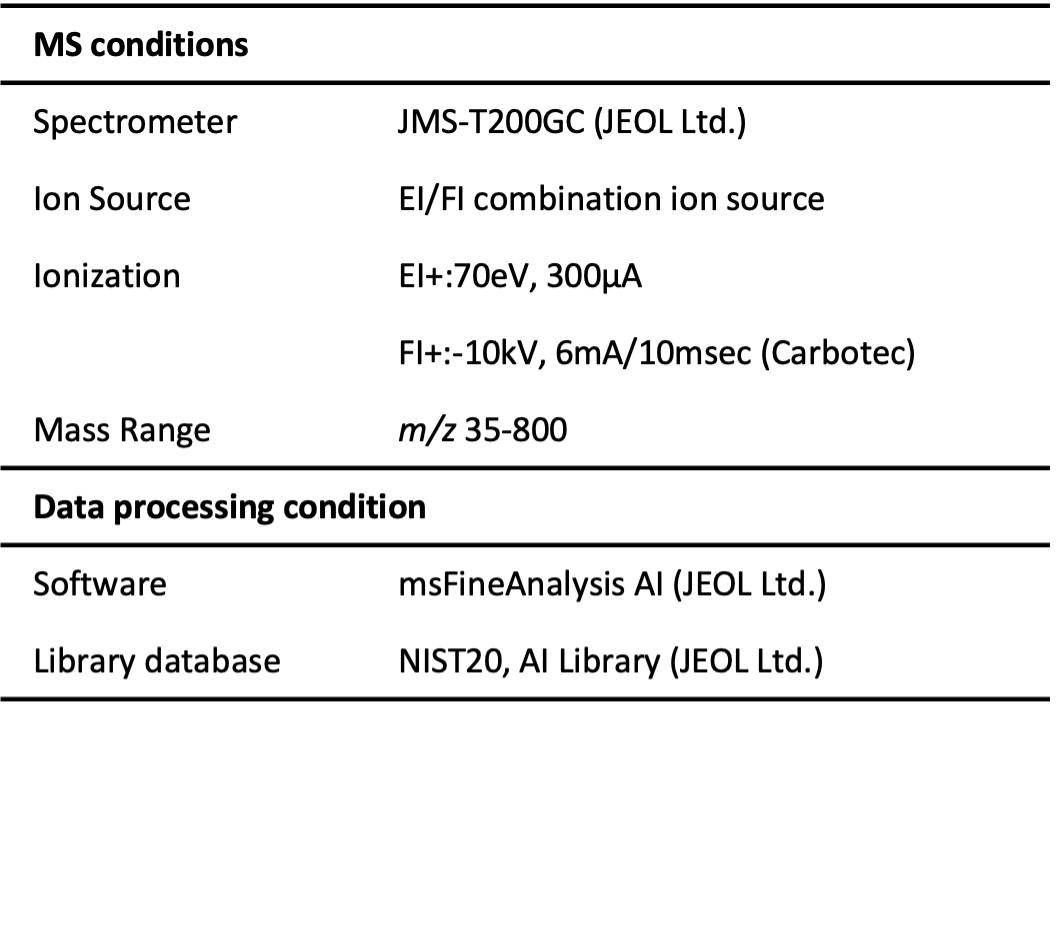
A commercially-available adhesive was used as a test sample in this study. We performed Py-GC-HRTOFMS measurements using both EI and field ionization (FI) modes with a combination EI/FI ion source. The qualitative data processing was performed with msFineAnalysis AI (JEOL). Measurement conditions are shown in Table 1.
Table 1 Measurement and analysis conditions
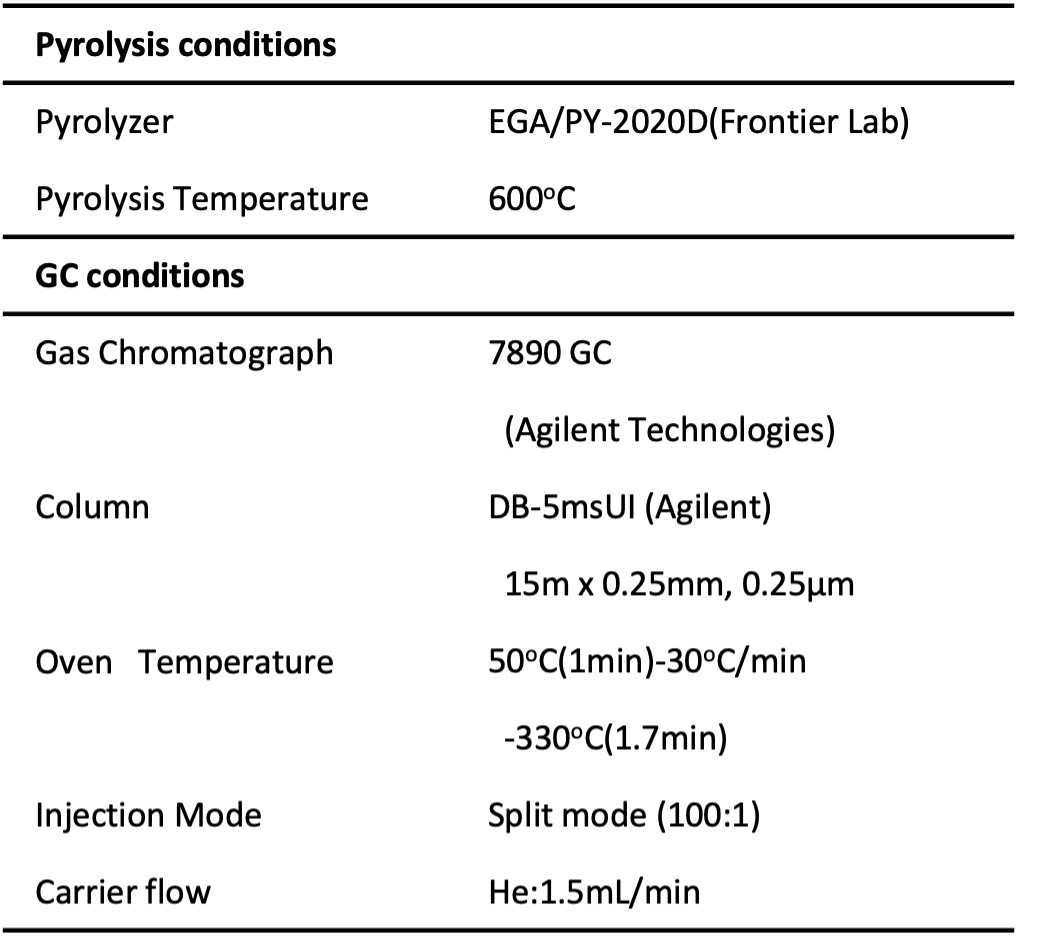

Results and Discussion
Extraction of off-flavor component by group analysis
Figure 1 shows an example of off-flavor compounds analysis result using group analysis function with "off-flavor list".
The chromatographic view on the left top shows the GC/EI data with the TICC marked by a solid black line. The bottom left view shows the soft ionization data with the TICC marked by a solid green line. The view on the right shows "Ion List" (applied the off-flavor list) detected in the analysis data. This off-flavor list contains 33 molecular formulas. If a compositional formula registered in this off-flavor list is present in the analysis data, the compound name of the off-flavor component, the type of odor, and the CAS number will be displayed in the Description column. In this study, multiple ions were detected that matched the molecular formula of the off-flavor component. This time, we focused on ion that match acetophenone (Sweet pungent odor, CAS No. 98-86-2) with the molecular formula C8H8O.
The blue peaks in each data of Figure1 represent the components containing C8H8O+ extracted from the chromatographic deconvolution result. The operator can select an ion such as C8H8O+ from the table and click the OK button at the bottom right of the GUI to immediately create a C8H8O tab, thus allowing for extraction of the components containing C8H8O+ (Figure 2).
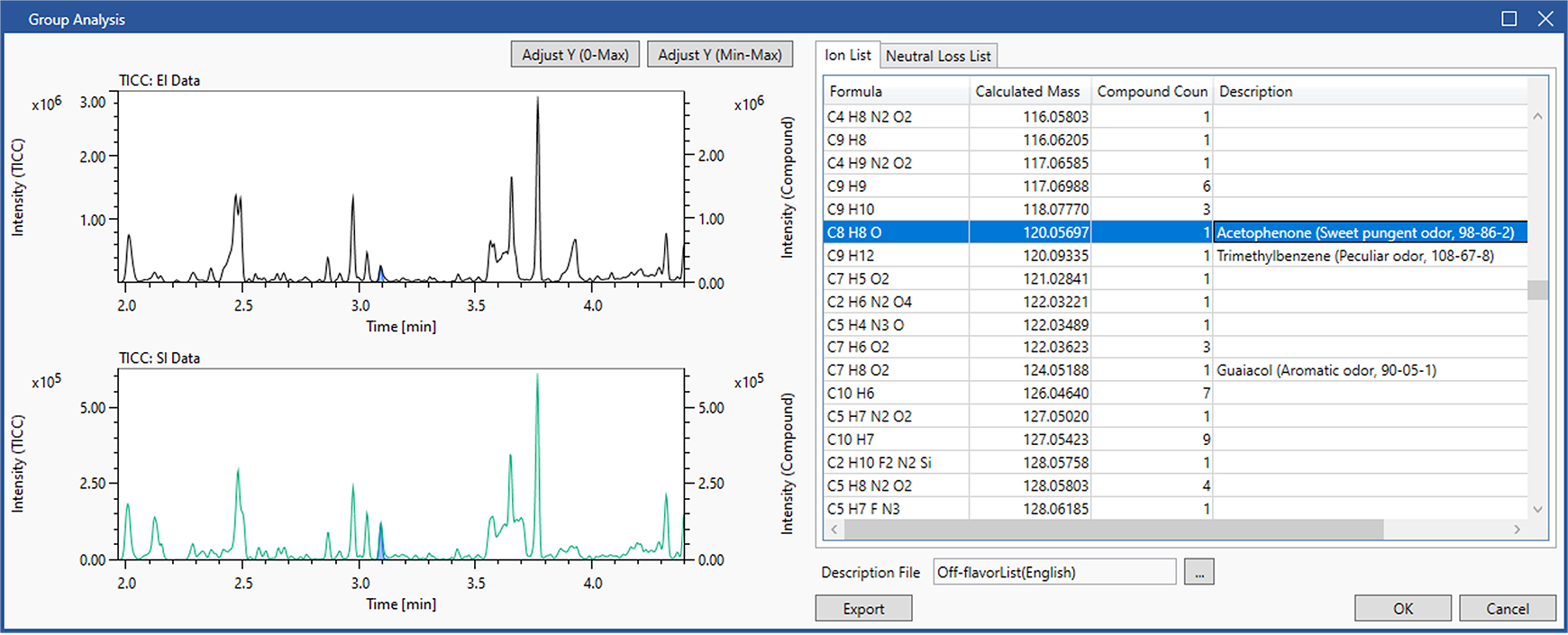
Figure 1 Group analysis result using off-flavor list
Figure 2 shows the extracted results for components containing C8H8O+ The Group Analysis function displays an "All" tab for the entire analysis results and up to 5 tabs for groups created for ions or neutral losses specified from the exact mass list in Figure 1. The ID and integrated analysis results are then shared between tabs. A total of 60 component peaks were observed in the analysis results, but it was possible to immediately extract only the peak containing C8H8O+. Detailed qualitative analysis results for this component are described in the next section.
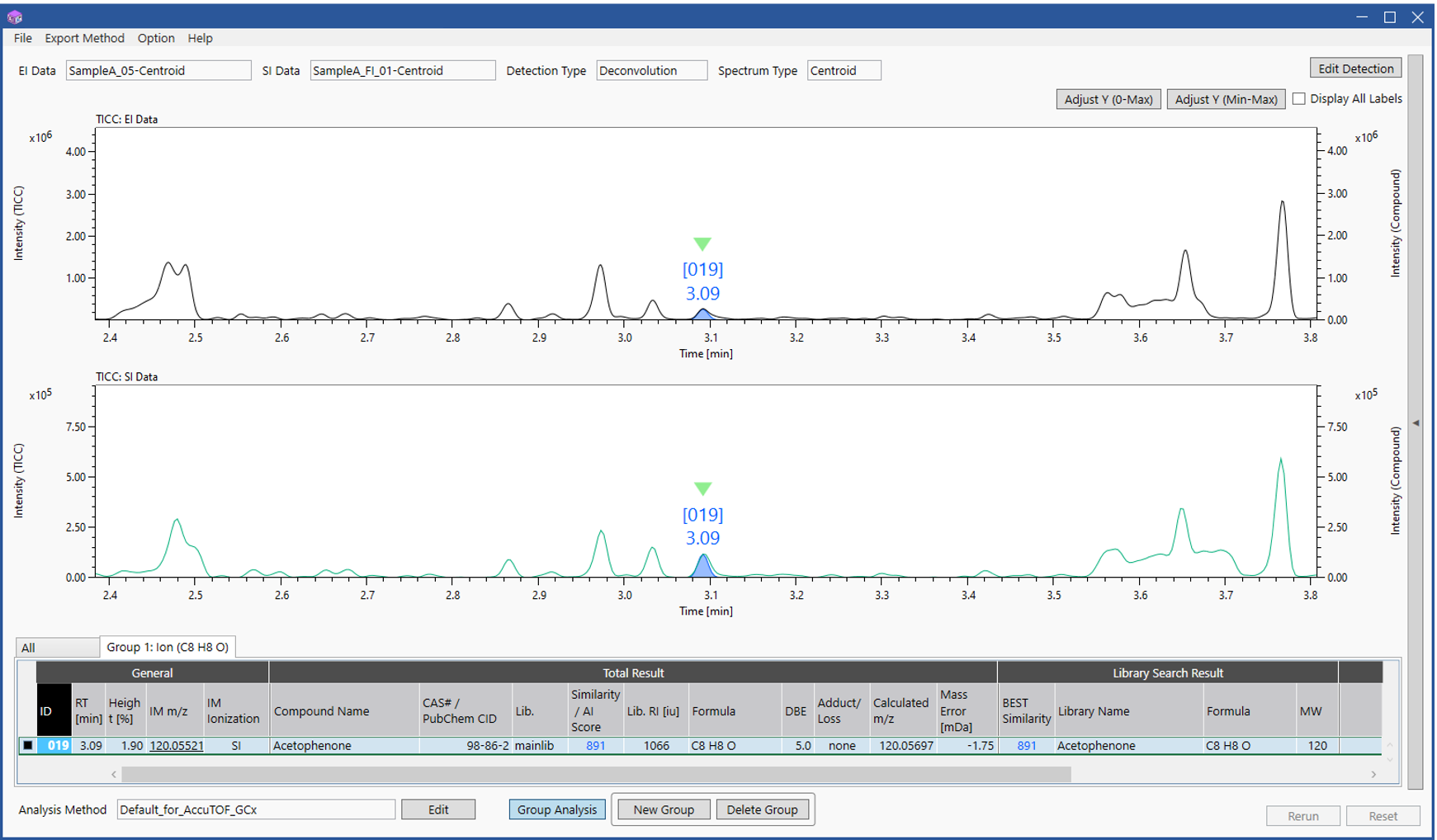
Figure 2 Group analysis result for C8H8O ion
Qualitative analysis of off-flavor component
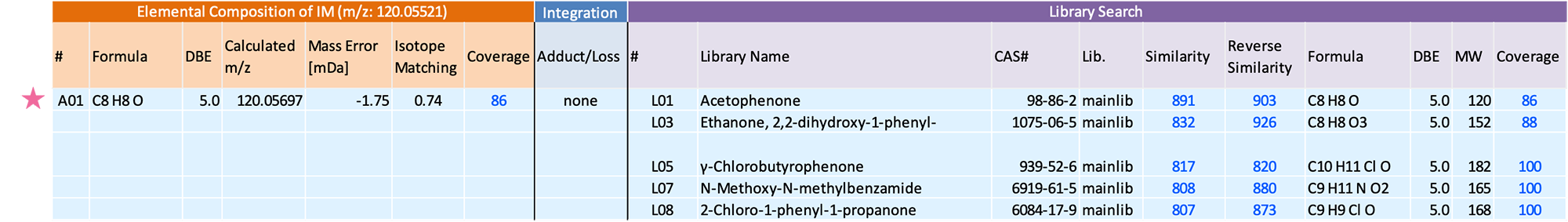
Figure 3 shows the mass spectra of the component extracted by group analysis function. m/z 120, which is presumed to be the molecular ion, was detected in both the EI and FI mass spectra, and the FI mass spectrum showed the molecular ion as the base peak. The integrated qualitative analysis result list (top 5 candidates) by msFineAnalysis AI is shown in Table 2 In the NIST Library DB search result, several highly similar compounds were presented. However, based on the estimated elemental composition of the molecular ion, the molecular formula was calculated to be C8H8O. Therefore, this component was presumed to be "Acetophenone" (Figure 4). From the above results, it was confirmed that off-flavor component was detected from the commercially available adhesive measured this time.
Even in an analysis such as pyrolysis GC-MS analysis in which many peaks are detected, off-flavor-derived peak could be extracted immediately.
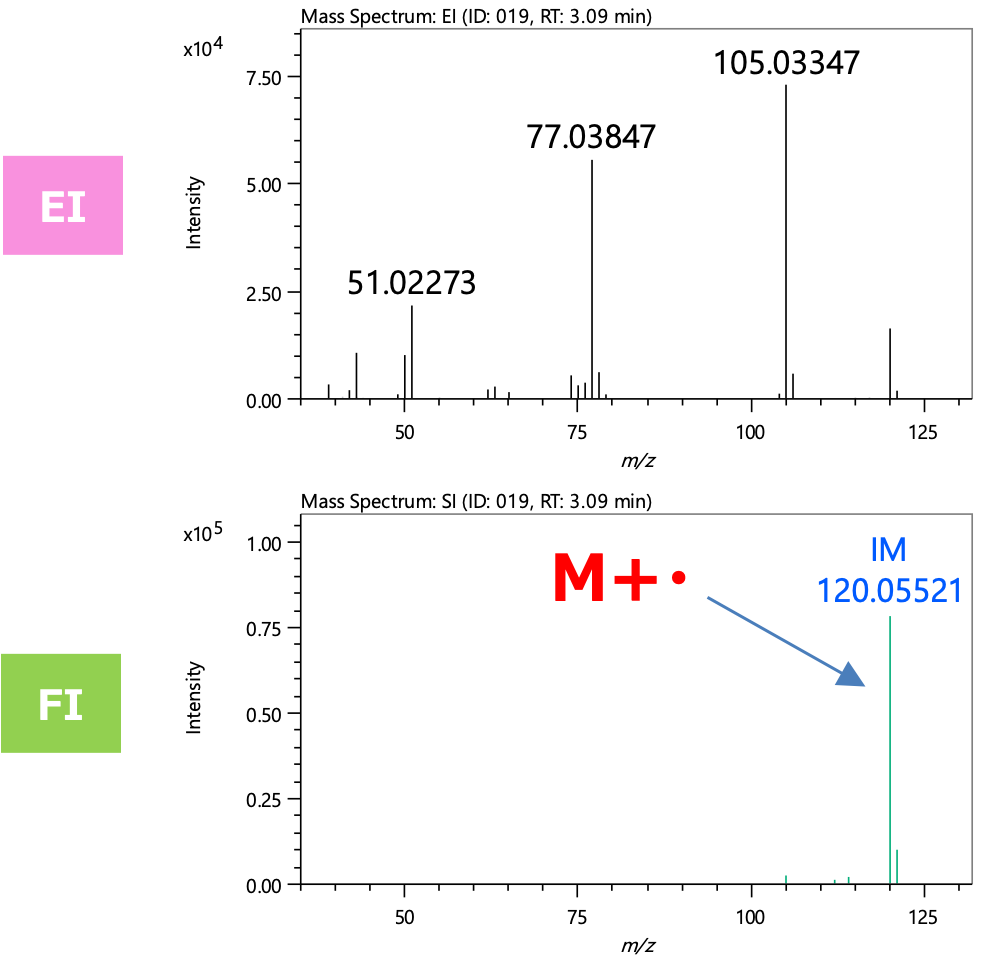
Figure 3 Mass spectra
Figure 4 Structural formula of Acetophenone
Table 2 Integrated qualitative analysis result using the msFineAnalysis AI
Conclusion
In this MSTips, we introduced an example of extracting off-flavor component in non-target analysis using the group analysis function of msFineAnalysis AI. By using the group analysis function, target analysis-like analysis can be performed even in non-target analysis, so detailed analysis in a shorter time can be expected.
Reference
- M. Ubukata, A. Kubo, K. Nagatomo, T. Hizume, H. Ishioka, A. J. Dane, R. B. Cody, Y. Ueda. Integrated qualitative analysis of polymer sample by pyrolysis–gas chromatography combined with high-resolution mass spectrometry: Using accurate mass measurement results from both electron ionization and soft ionization. Rapid Commun Mass Spectrom. 2020; 34:e8820.
- A. Kubo , A. Kubota, H. Ishioka, T. Hizume, M. Ubukata, K. Nagatomo, T. Satoh, M. Yoshida, F. Uematsu. Construction of a mass spectrum library containing predicted electron ionization mass spectra prepared using a machine learning model and the development of an efficient search method. Mass Spectrometry. 2023; 12: A0120.
Solutions by field
Related products
Are you a medical professional or personnel engaged in medical care?
No
Please be reminded that these pages are not intended to provide the general public with information about the products.


