Introduction of AI Structure Analysis Function in Automatic Structure Analysis Software msFineAnalysis AI
MSTips No. 388
Introduction
Electron ionization (EI) is one of the most popular ionization methods used in gas chromatography-mass spectrometry (GC-MS). Consequently, compounds are typically identified by a mass spectral database search using EI mass spectra. Because molecular ions are often weak or absent in 70 eV EI mass spectra, identification of unknowns can be difficult by EI alone. In these cases, soft ionization (SI) can be very helpful for producing and identifying molecular ions. Recently, JEOL began developing an integrated qualitative analysis workflow that automatically combines and interprets the information from EI and SI data. And then in 2018, we introduced our integrated qualitative analysis software "msFineAnalysis" which uses both EI and SI data to improve compound identification for GC-MS applications.
Despite the fact that msFineAnalysis was automatically able to determine the molecular formula and partial structure information from EI fragment ion formulas, the actual structural formulas still required manual analysis using chemical compositions. To address this, we then developed an automated structure analysis software package entitled "msFineAnalysis AI" which uses artificial intelligence (AI) to predict EI mass spectra from chemical structures. We have used our newly-developed AI model to create a database of predicted EI mass spectra for around 100 million compounds. In this work, we introduce AI structure analysis function in automatic structure analysis software msFineAnalysis AI.
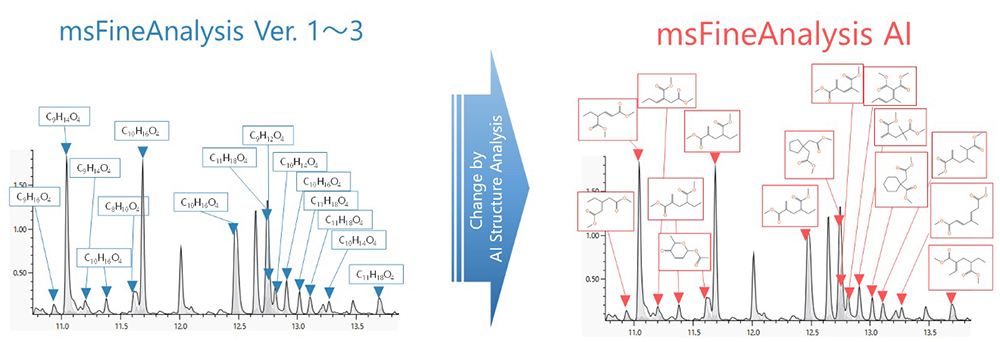
Figure 1 Image of analysis result in msFineAnalysis AI
About AI Structure Analysis Function
AI structure analysis function performs automatic structure analysis for unknown compounds using two AIs (main AI, support AI) that complementarily combine machine learning and deep learning.
Figure 2 shows the workflow of AI structural analysis by the main AI. In the main AI, a model for EI mass spectra prediction from structural formulas was constructed using deep learning, and predicted EI mass spectra of 100 million compounds were included in the software as an "AI library" database. The database search function using the "AI library" is implemented similarly to traditional library searches using the commercially available EI mass spectra database. Structural formula candidates are narrowed down by molecular formulas uniquely determined by integrated qualitative analysis, so more correct structural formulas can be obtained quickly. The predicted EI mass spectra were compared with measured EI mass spectra, then the scores were calculated from the spectral patterns, and candidate structural formulas were arranged in order of highest score. Finally, the correct structural formula is selected by combining the obtained structural formula candidates with the sample information and the knowledge and know-how obtained from the previous analysis.
Figure 3 shows the workflow of partial structure prediction by the support AI. The support AI assists interpreting analysis results by predicting the partial structure from the measured EI mass spectrum. It is possible to analyze the composition formula of fragment ions and neutral losses obtained from accurate mass analysis and assist in the interpretation of structural information proposed by the main AI.
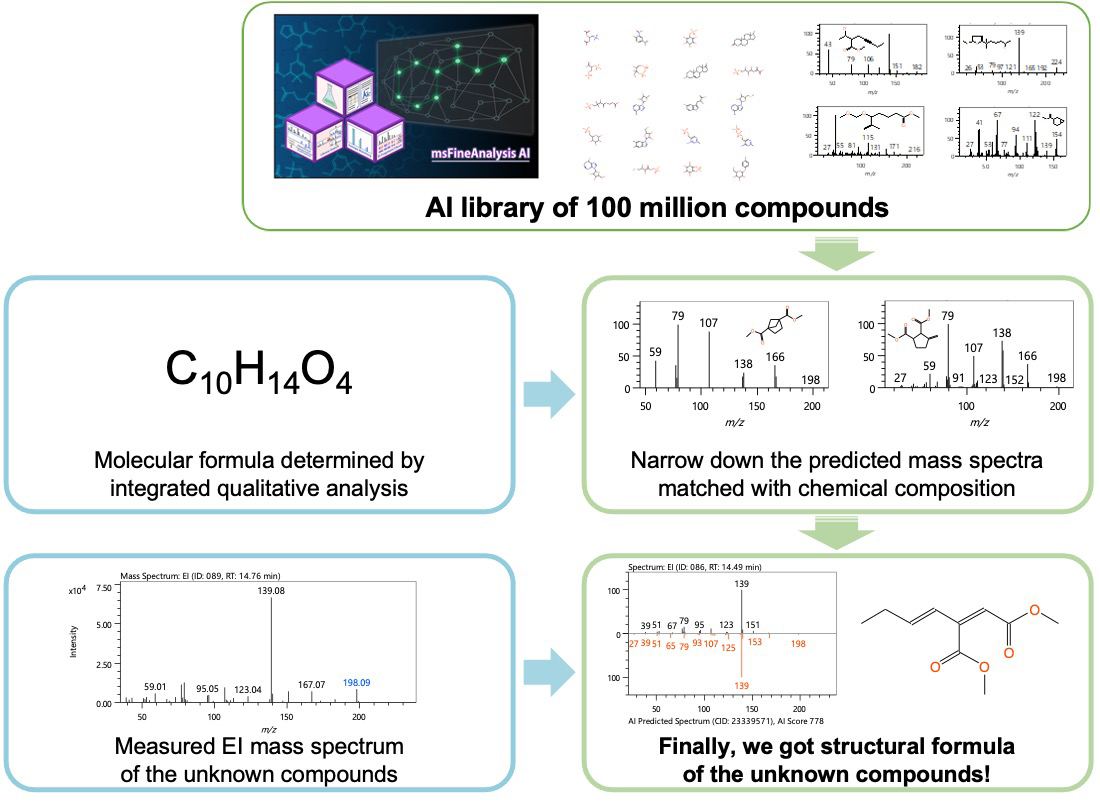
Figure 2 Main AI workflow
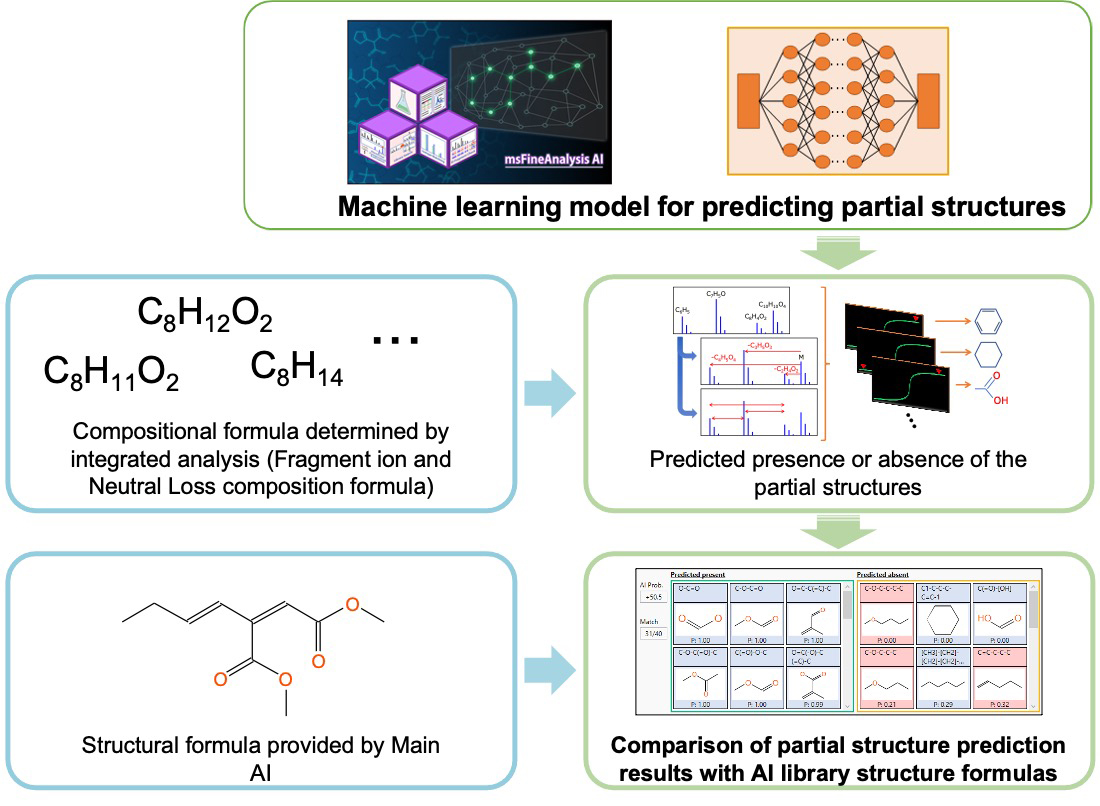
Figure 3 Support AI workflow
GUI of AI Structure Analysis Result
Figure 4 shows the AI structure analysis result of Acrylic Resin Oligomers by msFineAnalysis AI. The target of analysis is a dimer component that is not registered in the NIST library database. The left side of the analysis result screen shows the structure candidates by the main AI, and the right side shows the analysis results by the support AI. Detailed structural information can be obtained even for unknown compounds that have not been registered in the database.
On the main AI analysis result screen, a list of predicted structural formulas is shown at the bottom of the screen, and it is possible to check the AI structural analysis results all at once. The AI score indicates the similarity between the AI library and the measured mass spectrum, and it is shown at the bottom of each structural formula. Furthermore, information on the selected structural formula is posted at the top of the screen. We can see where the selected structural formula is in the histogram. It also includes a filtering function by partial structure and monomer, which enables structural analysis results to reflect the presence or absence of substructures predicted by the support AI described below.
On the support AI analysis result screen, predicted partial structure information is shown at the bottom of the screen. On the list, the left side is the partial structure predicted to be present, and the right side is the partial structure predicted not to be present. The partial structure with blue background matches the structural formula selected in the main AI, while the partial structure with red background does not match. Measured mass spectrum and the predicted composition formula of each fragment ion/neutral loss is posted at the top of the screen. It is also possible to confirm and edit comments for each estimated composition formula.
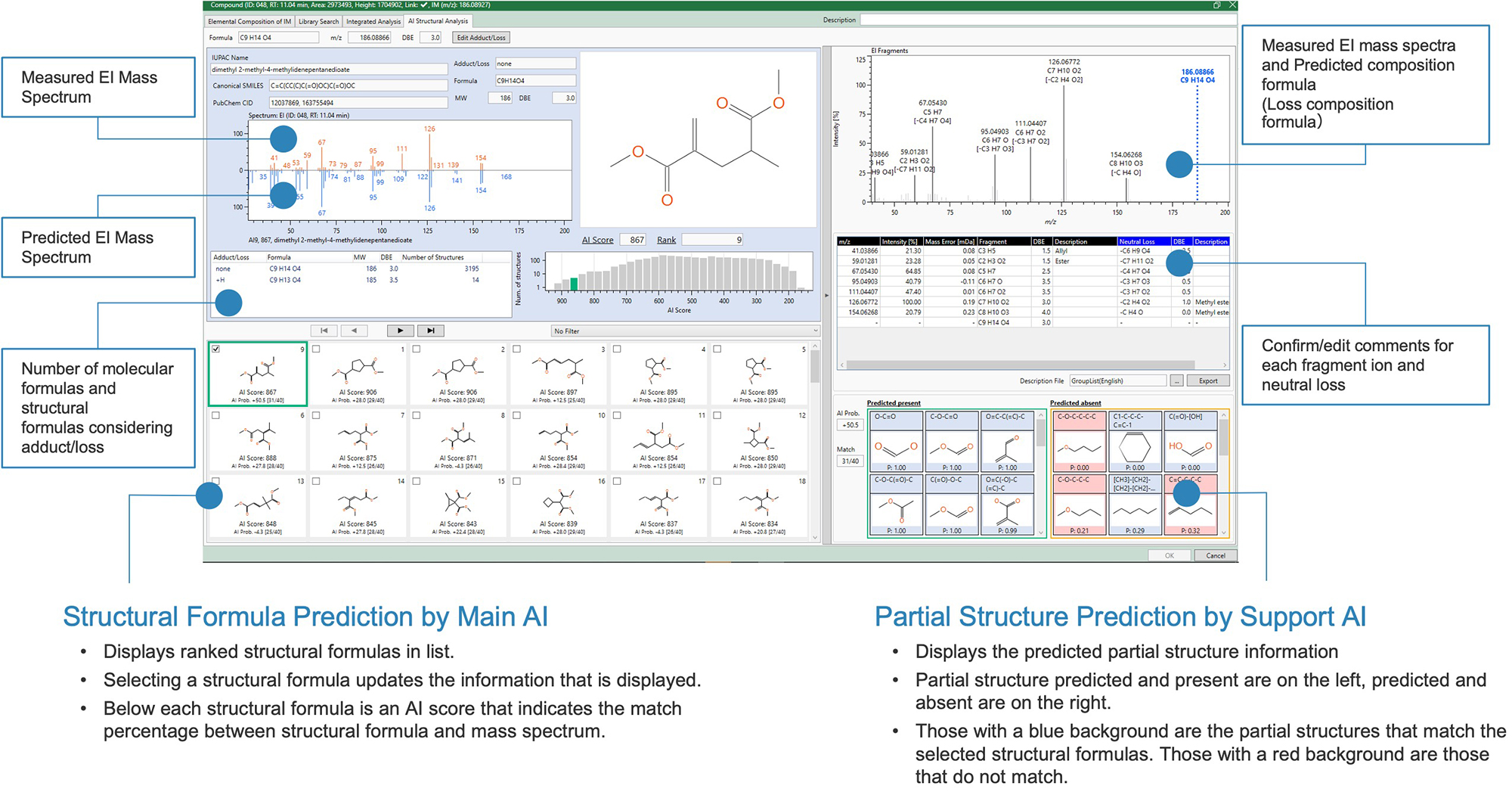
Figure 4 GUI of msFineAnalysis AI
Conclusion
In this MSTips, we introduced our newly-developed software msFineAnalysis AI, which contains AI structural analysis functionality to enhance qualitative analysis workflow. This software performs automatic structure analysis for unknown compounds using two AIs (main AI, support AI) that complementarily combine machine learning and deep learning. No knowledge of mass spectrometry and AI are required as the software automatically interprets complex mass spectra.
Qualitative analysis of GC-MS data can be greatly assisted by using EI and SI data together with msFineAnalysis AI, especially when trying to identify unknown compounds in complex samples.
Solutions by field
Related products
Are you a medical professional or personnel engaged in medical care?
No
Please be reminded that these pages are not intended to provide the general public with information about the products.


